An Approach to Fungal Diagnostics in Solid Organ Transplantation
David C. Gaston * ![]() , Marwan M. Azar
, Marwan M. Azar ![]()
- Department of Medicine, Section of Infectious Diseases, Yale University School of Medicine, New Haven, Connecticut, USA
* Correspondence: David C. Gaston![]()
Academic Editor: Maricar Malinis
Special Issue: Diagnosis and Management of Infections in Solid Organ Transplant Recipients
Received: November 14, 2018 | Accepted: December 24, 2018 | Published: December 29, 2018
OBM Transplantation 2018, Volume 2, Issue 4 doi:10.21926/obm.transplant.1804037
Recommended citation: Gaston DC, Azar MM. An Approach to Fungal Diagnostics in Solid Organ Transplantation. OBM Transplantation 2018; 2(4): 037; doi:10.21926/obm.transplant.1804037.
© 2018 by the authors. This is an open access article distributed under the conditions of the Creative Commons by Attribution License, which permits unrestricted use, distribution, and reproduction in any medium or format, provided the original work is correctly cited.
Abstract
The approach to diagnosing fungal infections following solid organ transplantation (SOT) is patient-specific. An assessment of an individual’s risk for particular infections, also understood as the pre-test probability, should guide appropriate diagnostic testing whereas diagnostic stewardship is needed to produce interpretable, actionable and cost-sensitive results. This review provides a cognitive framework for practitioners aiming to diagnose fungal infections in recipients of SOT, taking into consideration the benefits and limitations of currently available diagnostic platforms. In order to better characterize diagnostic utility for this population, we focus on clinical studies that include patients with SOT in full or in part. The goal of this review is to improve the diagnosis of fungal infections in patients with SOT by encouraging a thoughtful, systematic diagnostic approach.
Keywords
Solid organ transplantation; invasive fungal infection; clinical microbiology; diagnostic stewardship
1. Introduction
1.1 General Introduction
Transplantation of solid organs (including heart, lung, kidney, liver, pancreas, and small intestine) provides improved quality of life and prolonged survivorship in patients with end-stage organ dysfunction. Patients receiving transplanted solid organs are at increased risk for infectious complications, in great part due to intense systemic immunosuppression employed to prevent organ rejection. The risk for acquisition of specific infectious pathogens is dependent on the time elapsed since the day of organ transplantation, the net state of immunosuppression, surgical factors (prolonged operations, high transfusion requirement, technique of anastomosis, vascular thrombosis), any occurrence of organ rejection, and presence of indwelling catheters, among others (Figure 1) [1,2,3]. Although bacterial and viral infections are the most common infectious complications, invasive fungal infections in patients with transplanted solid organs carry high morbidity, including graft failure and loss, as well as increased mortality [1,4,5,6,7,8,9,10]. A high index of suspicion for invasive fungal disease must be maintained for early detection while balancing diagnostic and antifungal stewardship. This review aims to provide a practical reference for the practicing clinician when fungal infection is suspected in a patient with a transplanted solid organ.
The most commonly encountered fungal pathogens in the United States for SOT patients are Candida species (C. albicans, C. glabrata, and C. parapsilosis, less commonly C. krusei and C. tropicalis). Candida infections are most common in liver transplant recipients, followed by kidney, kidney-pancreas, and then lung transplant recipients [6]. Aspergillus species and Cryptococcus species are the next most common invasive fungi in this population [11,12]. Endemic mycoses (Histoplasma capsulatum, Blastomyces dermatitis, and Coccidiodes immitis / posadasii) are less common overall, with the risk impacted primarily by geographic exposure [9,13,14]. Mucorales infections are relatively uncommon, but carry a higher degree of morbidly and mortality given their fulminant clinical courses and limited available effective treatment options [10,15]. Other less common fungal infections, such as phaeohyphomycosis (caused by dematieacous or dark molds including Alternaria and Cladophialophora spp) or infections due to non-Aspergillus hyaline molds (such as Pseudallescheria and Fusarium spp), occur in the SOT with higher frequency than the immunocompetent population. Identification of these rare organisms is occurring with increasing frequency due to advances in clinical microbiology diagnostic technology [16,17,18].
1.2 Pre-Test Probability and Index of Suspicion
Patients with SOT are at a higher risk of invasive fungal infection as compared to the general population due to necessary immunosuppression [10,19]. The American Society of Transplantation Infectious Diseases Community of Practice largely advocates against routine screening for fungal infections in patients who have received SOT [20]. A notable exception is screening for invasive aspergillosis with Aspergillus galactomannan on bronchoalveolar lavage fluid in lung transplant recipients in centers with a high incidence of infection, as well as pre-transplantation screening (symptomatic, serologic, radiographic) for coccididiomycosis and blastomycosis in recipients from highly endemic areas. This recommendation does not extend to potential recipients in areas endemic for histoplasmosis because of a low likelihood of subsequent infection [21,22,23,24]. However, institutional standards may vary based on the risk assessment of the population served.
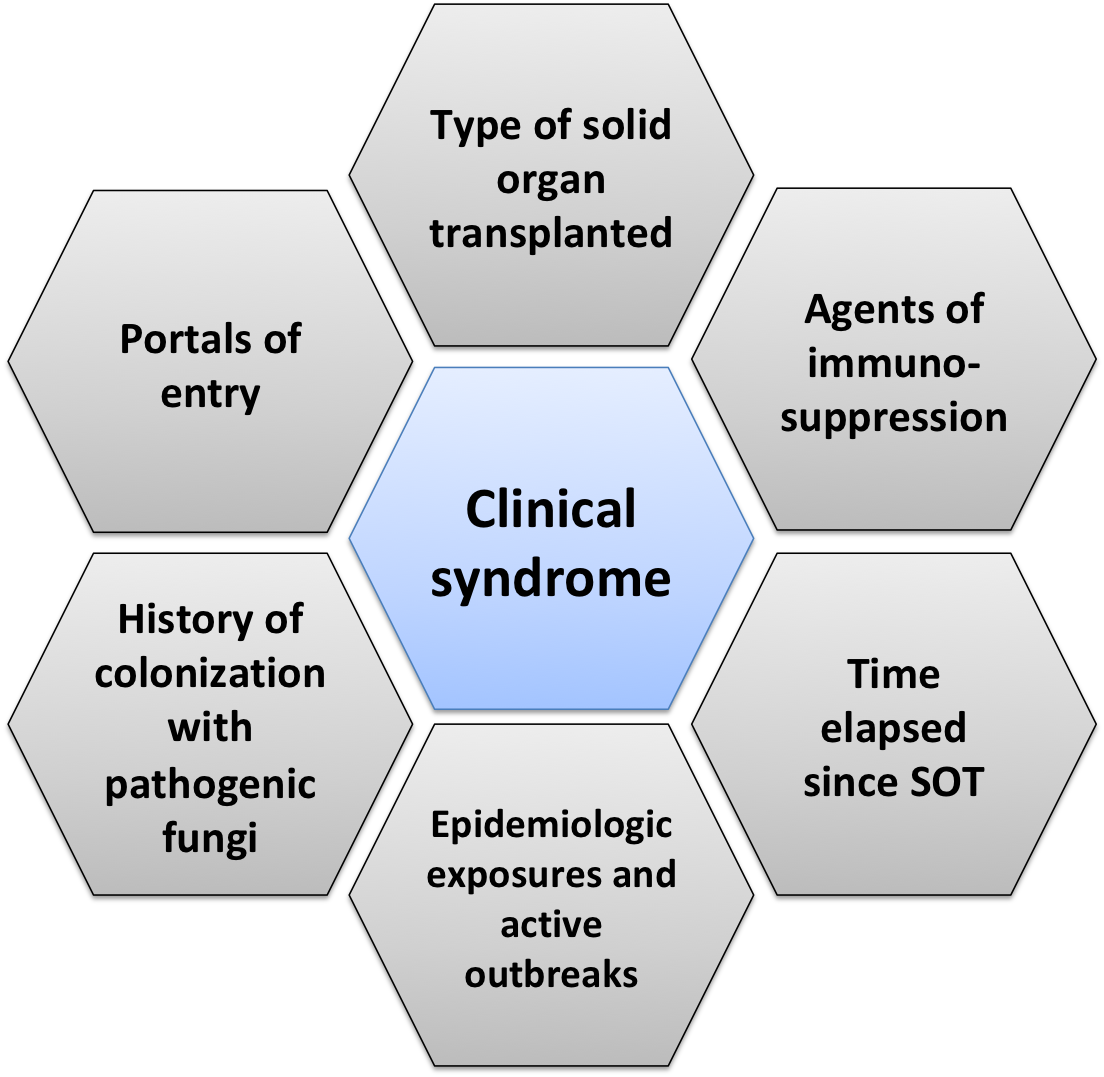
Figure 1 Drivers of pre-test probability for invasive fungal infections in Solid Organ Transplantation. Several factors drive the pre-test probability for invasive fungal infections in solid-organ transplantation (SOT). A. The most important is the presenting clinical syndrome, which is the constellation of historical, physical exam, laboratory and radiographic findings. B. Different transplanted organ types are associated with different risks based on the variable prevalence of IFI in those populations. C. The type of induction and maintenance immunosuppression impact the risk for specific IFI as does any intensification of immunosuppression. D. Several phases of post-transplant opportunistic infections have been defined as a function of time elapsed since SOT, each conferring risks for specific types of IFI based on immune cell recovery. E. Certain epidemiologic exposures are associated with increased risk for acquisition of fungi, especially endemic mycoses. Active outbreaks should be factored in to pre-test probability as well. F. Any history of colonization of pathogenic fungi increases risk of disease after immunosuppression G. Mucositis, presence of intravenous catheters and surgical complications can serve as portals for development of invasive infections especially with yeasts.
The first step in assessing the pre-test probability (and index of suspicion) for a particular fungal infection is determining its overall prevalence in a population of organ recipients. The transplanted organ is an important determinant in this assessment. For example, although Candida infection is the most common infection in SOT overall, infection with Aspergillus is most common in lung transplant recipients [25]. Evaluation of the PATH alliance registry in the US and Canada from 2004 – 2008 demonstrated patients with liver transplantation experienced the shortest time to invasive mold infections and had the highest rate of dissemination [19]. The next step is determining the risk (or incidence) of infection after a certain time post-organ transplantation. Following induction immunosuppression, innate and adaptive immunity are significantly impaired. While innate immunity recovers within weeks, adaptive immunity recovers much slower with T-cell immunity taking up to 1-2 years to reconstitute [26]. Several phases of post-transplant opportunistic infections have been defined, each conferring risks for specific types of fungal (and non-fungal) infections and all closely linked to the timing and type of immune cell recovery post-transplant [1]. Due to impaired neutrophil function combined with disrupted mucosal barriers and almost ubiquitous presence of vascular catheters in the first month post-transplantation, Candida spp are common fungal pathogens. Patients who develop prolonged neutropenia in the first month, particularly lung transplant recipients, are at increased risk for Aspergillus infections. Overall, Candida and Aspergillus species are the most commonly encountered pathogens in the first year after transplantation with Cryptococcus, endemic mycoses, and molds generally manifesting one year or later following transplantation [3,5,27].
Agents of induction and maintenance immunosuppression variably impact the risk of fungal infections [3]. Despite initial concerns for increased risk of fungal infection with Alemtuzumab, a monoclonal antibody to CD52 that targets both the cellular and humoral branches of the immune system, a similar incidence of fungal infections has been found as compared with other induction agents. When used as treatment for organ rejection, however, an increased incidence of fungal infections including aspergillosis, cryptococcosis and candidiasis was reported to occur within three months of therapy [28]. Similarly, the choice of maintenance immunosuppression has been associated with differing risks for invasive fungal infections (IFI). Mycophenolate has been associated with a lower risk for Pneumocystis jirovecii pneumonia while sirolimus has demonstrated in-vitro activity against Cryptococcus, Candida and Aspergillus [29]. As newer agents are added to the post-organ transplant armamentarium, additional risk assessments for associated IFI will be urgently needed in order to guide a diagnostic approach.
The distribution of fungi in the environment has led to the recognition of both large well-established environmental niches often associated with soil, terrain or vegetation as in the case of endemic fungi as well as micro-niches particular to specific transplantation centers in which certain fungal strains or resistance patterns predominate, often as a consequence of antimicrobial use practices. Person-to-person transmission of certain fungal infections, including P. jirovecii and C. auris, have occurred in transplant centers and any active outbreak should inform a particular patient’s pre-test probability [30,31].
Other important factors influencing the index of suspicion for IFI include concurrent medical conditions, prior infections or known colonization (including the microbiome of transplanted organ), and prior or current exposure to antifungal agents. Ultimately, and perhaps the most important determinant of the pre-test-probability, is the presenting clinical syndrome. This includes signs, symptoms, exposure history and characteristic radiographic findings (such as chest imaging for pulmonary aspergillosis, cryptococcosis, and invasive molds, or intracranial imaging for cryptococcal meningitis) [32,33].
1.3 Microbiology Testing Stewardship and Assay Interpretation
The diagnosis of an IFI in a patient with a transplanted solid organ occurs through careful assimilation of data from the clinical exam, radiography, and the microbiology laboratory. No individual diagnostic test is 100% sensitive and specific, and the prevalence of infection in the population studied influences positive and negative predictive values. Appropriate selection of diagnostic tests must be guided by the pre-test probability of a particular infection, itself based from multiple factors discussed in the previous section. Inappropriate or over-ordering of diagnostic tests carries the risk of misleading results with the possible consequences of antifungal overuse, increased costs, and worse outcomes.
An important concept in direct specimen testing is that available assays should only be performed on clinical specimens for which the test has been validated. This allows for adequate interpretation of results based on an understanding of performance characteristics. Performing testing on non-validated specimens should generally be discouraged as assay performance in these settings is uncharacterized and may be affected by specimen-laden interfering factors that can lead to false negative or false positive results. For samples not included in the manufacturer validation (regulated by the Food and Drug Administration) individual clinical microbiology laboratories can validate assays as Laboratory Developed Tests (LDTs). For tests that are developed or modified, Clinical and Laboratory Standards Institute Amendments (CLIA) mandates that individual laboratories establish the performance specifications for accuracy, precision, analytical sensitivity and specificity (including interfering substances), reportable range of results, reference intervals and test calibration. This system, which is overseen and enforced by CLIA regulators, has allowed LDTs to become an integral part of diagnostic testing, especially in immunocompromised populations where FDA-approved tests are lacking. Though extremely useful, it is important to recognize that the performance of LDTs may not be generalizable across transplant centers and are often based on fewer data than FDA-approved testing. Physicians should request these data from laboratories in order to better understand testing performance and how it applies to their patients.
Culture-based diagnostics have been well-established on essentially all clinical specimens. Fungal isolates are crucial for susceptibility testing and epidemiologic surveillance. In the setting of immunosuppression, particularly in the first year post-transplantation, serologic testing is often unreliable and negative serologic tests should not be used to exclude infections, including those related to fungi. Conversely, since fungal burden is higher, antigen-based testing is significantly more sensitive in this population. For example, the urine Histoplasma antigen was 93% sensitive in a series of solid-organ transplant recipients as opposed to 36% for serology [34].
As no one diagnostic test in the microbiology laboratory is perfectly accurate, and given the clinical complexity of patients with SOT, a certain degree of diagnostic uncertainty must be anticipated when attempting to diagnose an IFI. To attain a final diagnosis, multiple parallel methods of diagnosis must often be employed and rectified with the clinical syndrome. Available diagnostic testing with the highest sensitivity and specificity for a given suspected syndrome should be selected. The likelihood ratio for a given infection is influenced by the sensitivity and specificity of a chosen assay as well as the pre-test probability obtained from the overall index of suspicion. Overly-sensitive tests may lead to false positive results [35]. Certain causes of false-positive results have now been well established. These include false positive 1,3-β-D-glucan with heavy gauze exposure, gram-positive bacteremia, intravenous immune-globulin and beta-lactam-beta-lactamase antibiotics (although no longer common) [36]. Positive and negative results must be interpreted in context of each individual patient, recalling that many external factors affect the final test result. Considering the broader clinical syndrome is also necessary to differentiate an infectious pathogen from a colonizing organism, and certain clinical syndromes have not been convincingly linked with certain pathogens. For example, Candida pneumonia is thought to be exceedingly rare with few exceptions (tracheobronchitis in lung transplant recipients and hematogenous spread during intravascular infection) [37]. Additionally, methods based on isolate comparison to available databases, such as matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) are limited by the quality and currency of the reference standards. This vulnerability was exposed with the emergence of C. auris, an organism that is predictably misidentified or not identified by certain platforms, including earlier iterations of MALDI-TOF databases [38]. Updates to commercial identification systems will be required to ensure continued accuracy in diagnosing fungal infections. Finally, the method and quality of sample collection can significantly influence the reliability of a result. Methods of sample collection used for immunocompetent patients can be used in patients following SOT with similar diagnostic yield [39,40,41]. Collaborating with experienced specialists to select the optimal method of sample collection is essential to balance risks of invasive procedures with the possibility of inadequate sampling. The considerations listed above become clinically relevant in the context of low positive results, high clinical suspicion in the presence of negative results, or conflicting results.
2. Diagnostic Assays and Platforms for Fungal Infections in SOT
Diagnostic assays for fungal infections can be categorized as either culture dependent or culture independent, and broadly divided into closed and open platforms (Table 1). Closed platform assays are directed at a limited number of fungi. Examples of closed-platforms include assays detecting specific fungal components (1,3-β-D-glucan, galactomannan, Histoplasma antigen, Blastomyces antigen, and Cryptococcus antigen), serology, some polymerase chain reaction (PCR) based assays, and multiplex assays (FilmArray, T2Candida). These assays are developed to detect the most clinically significant fungi. More advanced platforms carry higher degrees of sensitivity and less cross-positivity, but organisms not included in the assay will not be detected. Conversely, open-system platforms analyze fungal genetic material or protein composition and compare results to a database for identification. These platforms include sequencing methods (metagenomic next generation sequencing) and matrix assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS). Such platforms aim to identify any fungi included in the reference database, and are limited only by the breadth and quality of the comparative database. A distinct advantage of these systems is the ability to expand and refine genetic or protein-based signatures, leading to an improved diagnostic ability. Notably, the most historically utilized open platform, and indeed the gold-standard of diagnosis, is direct morphologic identification of fungi from organisms cultured in the microbiology laboratory or observed in pathologic analysis of clinical specimens. The comparative reference in this case is the training and experience of the observer.
Table 1 Diagnostic methods for invasive fungal infection (IFI) in solid-organ transplantation (SOT).

2.1 Culture-Dependent Assays
Growth of yeast in blood cultures is often feasible, especially for Candida species, though the sensitivity is poor (~50%) [42]. Fusarium spp are unique among molds in their propensity to grow in blood cultures with a sensitivity of 48-82% in patients with invasive fusariosis [43,44]. Fungal blood cultures (Myco/F lytic) offer a modest advantage over conventional blood cultures (BACTEC) for mold identification (though likely not clinically significant) and none for yeasts [45]. Isolator Lysis-Centrifugation system-based culture is more sensitive for the diagnosis of endemic fungi, Cryptococcus and Malassezia [46]. Phenotypic identification of fungi in culture of clinical specimens is a longstanding standard of diagnosis. However, fungi, particularly molds and some endemic fungi, often grow slowly with some fungi (such as Histoplasma capsulatum) taking up to six weeks to form visible colonies. Grinding clinical specimens may prevent growth of Mucorales in culture because their aseptate hyphae are especially susceptible to disruption. Microbiology laboratories should be contacted when Mucorales are suspected to encourage mincing of tissue rather than crushing to enhance recovery of these fungi. Although morphologic growth characteristics can aid in identification, slow growth rates limit the clinical utility of fungal culture. Morphologic identification based on macroscopic colony characteristics and microscopic examination of reproductive structures (elicited by growth on nutrient-rich media such as cornmeal or potato dextrose agar) is often feasible down to genus level. However, further identification to species level is difficult and requires a high level of expertise in mycology, a skill that is unfortunately being lost as laboratories lose experienced technicians to time and automation. Fungal culture can be supplemented by nucleic acid probes, which are performed on colonies for confirmation. Most importantly, culture provides fungal isolates for susceptibility testing and genetic sequencing, crucial for monitoring resistance patterns in both clinical and epidemiologic settings [16].
2.1.1 MALDI-TOF MS. Matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) produces characteristic protein-derived spectra, which are compared to reference databases for rapid organism identification. MALDI-TOF MS is an open platform, limited only by the breadth of the reference databases. These databases are continuously updated and as such MALDI-TOF MS offers tremendous diagnostic potential. Presently, fungal growth in sub-culture to isolate individual colonies for analysis is necessary for adequate results [47]. Adding a solution of formic acid to the matrix-laden yeast colonies breaks down capsules and thick outer walls, improving the mass-spectral quality. MALDI-TOF MS has demonstrated high accuracy in the diagnosis of yeast and is being actively used for routine yeast diagnosis in clinical laboratories. Performance for molds has been highly variable, likely due to higher susceptibility to differences in processing methods, including culture conditions, extraction methods, hardware function and set thresholds. Notably, Mucorales have been the most difficult to identify with MALDI-TOF MS [48]. Overall, MALDI-TOF MS can provide correct identification to the level of species for fungi in over 90% of isolates, which is markedly higher than genetic based identification assays [48,49,50]. MALDI-TOF MS precision is likely to improve with inter-laboratory standardization of reagents, protocols, and instruments.
2.2 Culture Independent Assays
2.2.1 Histopathology. Morphologic identification of fungi from histopathologic specimens is a gold-standard of diagnosis. Histopathologic diagnosis with staining of tissue samples, an open platform, is the oldest method of identification and continues on most biopsy specimens obtained, is limited only by the knowledge and experience of the observer, and can provide rapid identification of fungi before growth in culture occurs [51]. The most useful stains for the diagnosis of fungi in tissue are the Gomori-methenamine silver and periodic acid Schiff stains which render black fungal forms on a green background and fuchsia forms on a pinkish background, respectively. When cryptococcosis is suspected, special stains such as the Fontana-Mason (highlights melanin) and mucicarmine (highlights capsule) stains can be utilized.
2.2.2 Antigen Detection. Antigen detection assays are available for polysaccharide antigens common to multiple fungi (1,3-β-D-glucan and galactomannan) as well as antigens specific for particular organisms (Blastomyces dermatitis, Histoplasma capsulatum, and Cryptococcus neoformans / gattii). Detection of pathogen-specific antigens in clinical samples must be interpreted within the entire clinical context as false positive results sometimes occur.
2.2.2.1 Galactomannan and β-D-Glucan. Galactomannan testing on serum or bronchoalveolar lavage (BAL) fluid is useful for the diagnosis of invasive aspergillosis. Only one assay is commercially available (Bio-Rad), and a systematic meta-analysis of fifty studies (including multiple categories of immunocompromised patients) determined the sensitivity ranged from 63-78% and specificity ranged from 85-93% based on the optical density threshold utilized [52]. Galactomannan antigen was positive in only one-third of patients with culture or biopsy-proven or probable invasive aspergillosis following SOT in Switzerland [4]. Similarly, less than half of patients with SOT and culture-identified aspergillosis had positive serum galactomannan testing [53]. When assessed for performance in a meta-analysis, the assay had a lower sensitivity and specificity in patients immunosuppressed due to SOT as compared to patients with hematologic malignancy [54]. Therefore, galactomannan testing should not be used to rule out invasive aspergillosis in SOT. Testing galactomannan on BAL is more sensitive than serum and may be particularly useful in lung transplantation. In a study of 16 lung transplant recipients, the sensitivity and specificity of galactomannan on BAL for diagnosing invasive aspergillosis with cutoff of ≥ 0.5 was 100% [55]. False positive results are reported in SOT patients with histoplasmosis (though interestingly Histoplasma antigen testing is rarely positive with aspergillosis) and with other factors including beta-lactam antibiotics, though the effect is increasingly rare [56,57,58]. The use of antifungal medication also decreases the sensitivity of detecting this antigen from serum [59]. Serial monitoring of serum galactomannan may be useful to monitor disease response in patients who have elevated levels [60].
Organisms that contain the polysaccharide 1,3-β-D-glucan in their cell wall may be detected by the 1,3-β-D-glucan assay in the setting of an IFI. The assay has poor sensitivity for organisms that contain little β-glucan in their cell wall including C. neoformans, Mucorales species and Blastomyces dermatitidis. Only one assay is commercially available (Fungitell, Associates of Cape Cod, Inc.), and is approved solely for serum samples. A meta-analysis of studies spanning 1995 to 2011 investigating the diagnostic utility of this assay in patients with and without organ transplants demonstrated a pooled sensitivity and specificity of 80% and specificity of 82%, respectively [61]. This study also suggested the greatest sensitivity and specificity for this test occurs with Pneumocystis jirovecii infection. Indeed this test may be most useful clinically when P. jirovecii pneumonia is suspected in a solid-organ transplant recipient, since GMS staining on BAL may be falsely negative [62]. In patients with lung transplantation, the assay demonstrated positive and negative predictive values that were too low to be useful for screening purposes [63]. Similarly, the assay demonstrated a sensitivity and specificity of 79.2% and 38.5%, respectively when used to evaluate BAL specimens from patients with lung transplants and proven or suspected IFI [64].
If a diagnosis is made, positive values can be trended over time to monitor therapy [65]. Although these assays are only FDA approved for serum samples (and BAL specimens for galactomannan), individual clinical microbiology laboratories can establish laboratory-developed tests to evaluate non-serum samples as well as to aid in the diagnosis of other fungi producing these polysaccharides. Such tests lack inter-laboratory standardization and must be rigorously validated but can be useful clinically. Testing 1,3-β-D-glucan on CSF samples was a highly useful adjunctive test for the diagnosis of iatrogenic fungal meningitis secondary to contaminated methylprednisolone acetate [66].
2.2.2.2 Histoplasma and Blastomyces Antigen. Histoplasma capsulatum antigen testing detects the presence of a specific cell wall polysaccharide and is primarily preformed on serum and urine samples. The sensitivity of urine Histoplasma antigen correlates with the burden of disease, with more severe infections associated with a higher frequency of antigen positivity [34,67]. Cautious interpretation of positive urine Histoplasma antigen must be taken as a high degree of cross-reactivity with other endemic fungi occur, particularly Blastomyces dermatitis [68]. Low positive results should not be quickly dismissed as false positives especially in endemic areas; in one study three quarters of low-positive results were clinically significant [69]. Histoplasma antigen can also be detected from CSF and BAL specimens, although similar interpretive caution is needed in patients with solid organ transplantation [68]. CSF Histoplasma antigen was found to be 81% sensitive in a series of 55 immunocompromised patients, 10 of which were recipients of transplantation [70].
An antigen assay for Blastomyces dermatitidis detects a cell wall polysaccharide and can be used for serum and urine samples. This assay is highly sensitive but similarly limited by a high degree of cross-reactivity, particularly with Histoplasma capsulatum and other fungal infections [71].
2.2.2.3 Cryptococcus Antigen. Polysaccharide antigens produced by Cryptococcus neoformans and Cryptococcus gattii can be detected by multiple commercially available assays using serum and CSF. These assays are highly sensitive (>90% in serum, >95% in CSF) and useful in the diagnosis of cryptococcal disease, including cryptococcal meningitis [72]. Initial high titers (>1:1024) serve as a poor prognosticator but titers do not respond to treatment in a sufficiently linear fashion to guide therapy [73]. A serum cryptococcal antigen titer greater than 1:64 is associated with a higher risk of cryptococcal meningitis in patients with SOT and cryptococcosis [74]. Additionally, isolated serum cryptococcal antigen titers ranging from 1:10 to 1:80 in the absence of other etiology for disease were adequate for chronic suppressive fluconazole in a case series of five patients with SOT [5].
2.2.3 Serology. Serology can help determine previous exposures and follow response to therapy in established infections, but positive and negative serologic testing must be interpreted in context of all other available data. Particular attention must be paid to geographic exposures with serologic testing for endemic mycoses. Serology for coccidioidomycosis continues to be the primary non-culture or pathology-based diagnostic method for this infection. A positive titer as low as 1:2 is considered clinically relevant and warrants treatment in patients with SOT [75]. Serologic criteria for acute histoplasmosis have been established by The Council of State and Territorial Epidemiologists [76]. Serology is not useful for the diagnosis of invasive aspergillosis and is mainly reserved for workup of allergic bronchopulmonary aspergillosis, a non-infectious allergic manifestation.
Interpretation of serologic testing poses a particular challenge in patients with SOT because immunosuppression dampens immune responses producing potentially false negative results. In addition, positive serology may represent acute or past infection and making the distinction can be difficult. Clinically informative serologic testing is available for histoplasmosis, coccidioidomycosis [77] and blastomycosis [78].
2.2.4 Polymerase Chain Reaction (PCR). Detection of fungal pathogens on direct clinical specimens utilizing polymerase chain reaction (PCR) based techniques commonly involves assay development in individual clinical microbiology laboratories with in-house validation. Certain FDA CLIA-certified laboratories have made such testing commercially available. Testing for fungi with LDT PCR assays can be highly sensitive. For example, a pooled meta-analysis of PCR based methods of detection of Candida species from blood samples estimated an overall sensitivity and specificity of 95% and 92%, respectively [79]. However, a lack of intra-laboratory standardization and clarity in assay performance characteristics renders interpretation of assay results difficult.
A pertinent example is the use of PCR to detect Aspergillus species from blood, respiratory, and tissue specimens. As no standardized method of detection is used across clinical microbiology laboratories, the sensitivity and specificity of individual platforms in differing laboratories can vary widely [80,81]. In a Cochrane meta-analysis that included patients with solid organ transplantation, the sensitivity and specificity of PCR for the diagnosis of invasive aspergillosis ranged from 58.0 to 80.5% and 78.5 to 95.2%, respectively with a mean sensitivity and specificity of 80.5% and 78.5% [82]. Notably, the sensitivity was estimated to decrease to 58.0% with an increase in the specificity to 96.2% with consecutive testing. More recent studies demonstrate improved PCR detection of Aspergillus in CSF samples, with 75% sensitivity and 98.3% specificity as well as positive and negative predictive values of 90% and 95.2% [83]. Similar trends are noted with PCR detection of other fungal infections, specifically P. jirovecii in patients with lung transplantation [84].
Alternatively, PCR can be applied to an unidentified fungal isolate. Fungal speciation can be achieved by primer binding to conserved internal transcribed spacer (ITS) regions in fungal ribosomal RNA, as well as other regions (such as elongation factor 1-alpha and beta tubulin) [50,85,86]. Fungi that do not contain primer-binding sites cannot be sequenced and therefore cannot be identified by this method. Additionally, sequencing data matching the organism in question must be present in the database used for comparison. These methods are similar to Sanger sequencing for identification but focus on a smaller portion of genetic material to provide identification. Sensitivity of broad-range PCR tests are lower than those directed at specific fungi.
2.2.5 Film Array. FilmArray (BioFire, Salt Lake City, UT / Biomerieux) is a multiplex-PCR-based closed platform that requires a growth-phase step in liquid culture. The FilmArray assay approved for serum samples detects multiple organisms, but the only fungal organisms detected are Candida species (C. albicans, C. tropicalis, C. glabrata, C. parasipolis, and C. krusei). The primary benefit of this assay is the rapidity of detection from positive cultures without the need for sub-culturing. This assay can be useful in quickly detecting common pathogens for which a given patient is at risk. In a study of liver transplant patients, a multiplex FilmArray did not miss infections detected by conventional methods [87]. Clinically important fungi not included on the panel were accordingly missed in multi-center trials comparing this assay to conventional diagnostic methods [88,89]. However, this assay had a sensitivity and specificity of 99.2% and 99.9%, respectively, for Candida species from clinical specimens, demonstrating the utility of this assay when considering the diagnostic time saved [88]. These studies highlight benefits of this rapid detection platform for common pathogens but emphasize that parallel methods are needed to detect pathogens not included in the assay. A separate FilmArray panel was investigated in a pre-clinical setting for CSF samples to detect cryptococcal meningitis, but was only 64.3% specific for Cryptococcus neoformans and gattii [90]. Commercially available FilmArray assays benefit from inter-laboratory standardization as compared to LDT PCR assays, which has implications for result interpretation for clinical care and research across institutions.
2.2.6 T2Candida. The T2Candida system (T2 Biosystems, Lexington, MA) is a closed platform that detects the five most clinically significant Candida species (C. albicans, C. tropicalis, C. krusei, C. parapsilosis, and C. glabrata). The assay, which is based on PCR amplification of Candida DNA followed by measurement of T2 magnetic resonance of DNA-probe complexes, detects genetic material directly from blood samples within 4 hours, with all processing and amplification steps contained within the device. In an initial trial comparing detection of candidemia from both patient-derived and spiked samples with conventional blood cultures, the overall sensitivity and specificity for the T2Candida system was 91.1% and 99.4%, respectively, a significant improvement from the sensitivity of the current gold standard of blood cultures (only ~50% sensitive for Candida species) [91,92]. A subsequent study that also included patients with solid organ transplants detected significantly more episodes of candidemia in patients receiving antifungal agents as compared to conventional culture methods [42]. This system may have utility for early diagnosis, treatment decisions, and possibly prognosis in multiple patients populations at risk for Candida bloodstream infections, including patients with solid organ transplants [93,94,95]. The primary benefit may be in antifungal stewardship rather than diagnosis given the current reagent cost of this system, but cost modeling suggests the system could yield savings [96,97].
However, the T2Candida system has several pitfalls [42]. Though the Mylokanis study reported a sensitivity above 90% overall, the true sensitivity is likely to be lower. Indeed, the majority of positive samples in the study were contrived specimens (250 of 257) and 44% of these contrived samples contained a yeast concentration (11-100 Candida CFU/mL) significantly higher than the mean concentration (1 CFU/mL) in initial blood cultures of candidemic patients [98]. This implies a diminished negative predictive value of a negative test in a patient with a high-pretest probability for candidemia and reduces the real-world utility of T2 in guiding initiation or continuation of antifungals in this setting. Additionally, this assay does not detect several clinically important agents of candidemia (such as C. kefyr, C. lusitanea, C. guilliermondii, and C. auris), which comprise around 5-10% of candida infections in SOT patients [1,12]. The assay is also prone to technical errors, yielding indeterminate or invalid results in >50% cases in the original study, potentially impacting workflow. No prospective clinical trials comparing conventional fungal diagnosis with this system and conventional methods have been conducted in SOT patients to determine if the time saved in diagnosis translates to reduced morbidity, mortality, or cost savings.
2.2.7 Next Generation Sequencing and Metagenomics. Next generation sequencing (NGS), or sequencing methods developed after Sanger sequencing, are PCR-based methods in which entire pathogen genomes can be sequenced in a relatively short time due to parallel high throughput amplification of small genetic sections. Metagenomics is the process of sequencing multiple organisms present in a single sample and frequently utilizes sequencing technology such as NGS. These are open platforms in which sequencing results are compared to reference databases to generate identifications. Next generation sequencing can be applied to culture or directly to clinical specimens. The clinical utility of NGS is currently being evaluated in SOT patients, and appears to be most useful for viral identification pediatric populations but recent studies suggest how these platforms may be used in the future [99]. For example, NGS facilitated identification of a nosocomial outbreak of P. jirovecii involving patients with SOT in France [100]. Additionally, a metagenomics approach identified P. jirovecii DNA in blood, BAL fluid, sputum, and tissue [101]. Although it can be difficult to differentiate colonizing organisms from those leading to disease, detection of pathogenic organisms by metagenomic NGS is possible [102]. Next-generation sequencing currently suffers from high cost and lower organism-specific sensitivity as compared to traditional PCR so providers should make every attempt to send pathogen-specific PCR testing based on pre-test probability.
3. Future Directions
Although microbiology laboratories often perform similar assays, reagents and methods vary, leading to lack of standardization and difficulty in inter-laboratory result interpretation. In the past decade, the WHO introduced testing standards for several transplant-associated viruses including CMV, EBV, and BKV designated in international units/ml. As a result, some improvement in variability was seen (especially with EBV testing) but significant inter-laboratories discrepancies still exist, underscoring the challenges associated with such endeavors [103]. Similar efforts can be made in standardizing platforms for IFI diagnosis.
Combining conventional methods to detect fungi increases diagnostic yield in SOT recipients by compensating for limited sensitivity and specificity of individual platforms [75,104,105]. Using microbiology assays with other diagnostic modalities, such as radiography, can also enhance diagnosis. For example, abnormal 18-FDG PET/CT results can increase the index of suspicion for IFI in patients with SOT and direct diagnostic evaluation [106,107]. Such methods should not be used in isolation but should rather be incorporated into algorithms to guide definitive testing for IFI.
Another approach to future research involve optimizing currently available methods for a more efficient means of detection. MALDI-TOF MS holds great promise in its ability to serve as a rapid open platform, but the necessity of fungal sub-culturing limits the utility even of this assay. Bacterial pathogens can be reliably detected by MALDI-TOF MS directly from positive blood cultures in immunosuppressed patients, including patients with SOT raising the possibility for direct identification of fungemia using MALDI-TOF MS [108].
Peptide nucleic acid fluorescent in situ hybridization (PNA-FISH) is another closed platform assay that can rapidly detect bloodstream infections of Candida species with similar performance to more widely utilized methods of detection [109]. In one study, this platform demonstrated a markedly higher rate of positive identification as compared to MALDI-TOF MS [110]. However, PNA-FISH has not been evaluated in the context of solid organ transplantation.
Rapid multiplex PCR assays have seen an expanded role in the diagnosis of fungal and bacterial infections in immunocompromised patients. However, commercially available systems often detect no fungal pathogens other than Cryptococcus on CSF [111,112]. PCR based assays incorporating fungi responsible for the majority of disease in SOT patients are currently under development and have been studied in patients with hematologic malignancies [113,114]. Additionally, multiplex FilmArray was investigated for direct analysis of clinical samples other than serum and CSF [115]. Research into the use of this assay for rapid diagnosis of Candida infections in SOT using samples other than serum or CSF, such as respiratory samples from patients with lung transplantation, could lead to improvements in both adequate treatment and optimal stewardship. Further utilization of these assays may significantly shorten time to diagnosis and enable the most appropriate antifungal regimens.
A current limitation to multiplex assays is the production of a binary result; an organism is either detected or it is not. These qualitative assays do not yield organism burden, which may be clinically useful. Although detection of some fungi almost always signifies infection (ex: Cryptococcus), detection of others (ex: Candida, P. jirovecii) may indicate colonization rather than pathogenicity. This distinction can often be made based on clinical presentation, but may not be always possible, especially in acutely ill patients. The high sensitivity of newly developed open and closed platforms may lead to more false positive results and more mis-diagnoses of IFI. This in turn can lead to overlooking of the true pathogenic process. Semi- and fully quantitative methods can determine the burden of organism present and potentially differentiate between colonization and pathogenicity [116].
Broadening the use of other non-multiplexed assays could enhance care of patients with organ transplants. Routine PCR can be adapted to challenging specimen types, as is the case of Aspergillus fumigatus DNA detected from formalin-fixed samples of a lung transplant recipient [117].
A notable gap in the study of SOT patients with IFI exists for the diagnosis of Mucorales infections. These are less common in patients with SOT as compared to hematologic transplantation, but invasive infection carries a higher level of mortality than other IFI. No rapid diagnostic methods are available, and current methods of identification (phenotypic identification and MALDI-TOF MS) rely on growth in culture. This is a significant barrier to diagnosis given the often poor yield of culture. Methods enhancing the diagnosis of Mucorales infections are a significant need in this field.
Novel point-of-care methods of invasive fungal detection are under development and clinical testing. An assay utilizing thermal desorption-gas chromatography/mass spectrometry determined characteristic signatures of volatile metabolites exhaled from patients with invasive aspergillosis, including patients with SOT [118]. This assay represents an open platform potentially capable of detecting volatile metabolites from other pathogens. Adaptation of this platform to provide early detection for Mucorales infections is a promising prospect. A separate point-of-care assay utilizing a lateral-flow device demonstrated sensitivity and specificity of 91% and 83%, respectively, for pulmonary aspergillosis when BAL fluid from SOT recipients was directly applied to the testing platform [119].
As new organ-types become transplanted with increasing frequency, data is emerging regarding IFI complicating these transplantations. For example, small intestine transplantation carries a similar risk for IFI as other transplanted organs, with Candida and Aspergillus infections occurring most commonly [120]. Additional research is needed into the risk factors associated with IFI in emerging procedures such as vascularized composite allotransplantation as well as optimal methods of early detection based on the epidemiology of infection.
4. Summary
The primary challenge in the future of diagnostic testing for fungal infections is the correct application of available testing in an efficient and economic practice. The clinical microbiology laboratory in any transplant center is a vital resource in the care of patients with SOT. Index of suspicion for IFI must be maintained at an appropriate level based upon the individual patient characteristics and the broader environmental context of the patient and transplant center. Microbiology stewardship must be employed to avoid misleading results and incurring unnecessary costs. Result interpretation must be predicated upon a basic understanding of the mechanisms and limitations of a given testing modality and squarely based in the varying risks and exposures for an individual patient. The clinical microbiology laboratory resides at the center of these issues. Further work towards optimizing current platforms and developing future standards of diagnosis will continue the forward progression of the transplant infectious diseases field.
Author Contributions
David Gaston composed, edited, and approved the manuscript; Marwan Azar edited and approved manuscript as well as composed the table and figure.
Competing Interests
The authors have declared that no competing interests exist.
References
- Fishman JA. Infection in organ transplantation. Am J Transplant. 1st ed. 2017; 17: 856-879.
- Silveira FP, Kusne S, AST Infectious Diseases Community of Practice. Candida infections in solid organ transplantation. Am J Transplant. 2013; 13 Suppl 4: 220-227. [CrossRef]
- Anesi JA, Baddley JW. Approach to the Solid Organ Transplant Patient with Suspected Fungal Infection. Infect Dis Clin North Am. 2016; 30: 277-296. [CrossRef]
- Neofytos D, Chatzis O, Nasioudis D, Boely Janke E, Doco Lecompte T, Garzoni C, et al. Epidemiology, risk factors and outcomes of invasive aspergillosis in solid organ transplant recipients in the Swiss Transplant Cohort Study. Transpl Infect Dis. 2018; 20: e12898. [CrossRef]
- Wald-Dickler N, She R, Blodget E. Cryptococcal disease in the solid organ transplant setting: review of clinical aspects with a discussion of asymptomatic cryptococcal antigenemia. Curr Opin Organ Transplant. 2017; 22: 307-313. [CrossRef]
- Andes DR, Safdar N, Baddley JW, Alexander B, Brumble L, Freifeld A, et al. The epidemiology and outcomes of invasive Candida infections among organ transplant recipients in the United States: results of the Transplant-Associated Infection Surveillance Network (TRANSNET). Transpl Infect Dis. 2016; 18: 921-931. [CrossRef]
- Kritikos A, Manuel O. Bloodstream infections after solid-organ transplantation. Virulence. Taylor & Francis; 2016; 7: 329-340. [CrossRef]
- Klingspor L, Saaedi B, Ljungman P, Szakos A. Epidemiology and outcomes of patients with invasive mould infections: a retrospective observational study from a single centre (2005-2009). Mycoses. 2015; 58: 470-477. [CrossRef]
- Kauffman CA, Freifeld AG, Andes DR, Baddley JW, Herwaldt L, Walker RC, et al. Endemic fungal infections in solid organ and hematopoietic cell transplant recipients enrolled in the Transplant-Associated Infection Surveillance Network (TRANSNET). Transpl Infect Dis. 3rd ed. 2014; 16: 213-224.
- Pappas PG, Alexander BD, Andes DR, Hadley S, Kauffman CA, Freifeld A, et al. Invasive fungal infections among organ transplant recipients: results of the Transplant-Associated Infection Surveillance Network (TRANSNET). Clin Infect Dis. 2010; 50: 1101-1111. [CrossRef]
- Taimur S. Yeast infections in solid organ transplantation. Infect Dis Clin North Am. 2018; 32: 651-666. [CrossRef]
- Neofytos D, Fishman JA, Horn D, Anaissie E, Chang C-H, Olyaei A, et al. Epidemiology and outcome of invasive fungal infections in solid organ transplant recipients. Transpl Infect Dis. 2010; 12: 220-229. [CrossRef]
- Nel JS, Bartelt LA, van Duin D, Lachiewicz AM. Endemic mycoses in solid organ transplant recipients. Infect Dis Clin North Am. 2018; 32: 667-685. [CrossRef]
- Osawa R, Alexander BD, Forrest GN, Lyon GM, Somani J, del Busto R, et al. Geographic differences in disease expression of cryptococcosis in solid organ transplant recipients in the United States. Ann Transplant. 2010; 15: 77-83.
- Cuenca-Estrella M, Bernal-Martinez L, Isla G, Gomez-Lopez A, Alcazar-Fuoli L, Buitrago MJ. Incidence of zygomycosis in transplant recipients. Clin Microbiol Infect. 2009; 15 Suppl 5: 37-40. [CrossRef]
- McCarty TP, Baddley JW, Walsh TJ, Alexander BD, Kontoyiannis DP, Perl TM, et al. Phaeohyphomycosis in transplant recipients: Results from the Transplant Associated Infection Surveillance Network (TRANSNET). Med Mycol. 2015; 53: 440-446. [CrossRef]
- Sitterlé E, Giraud S, Leto J, Bouchara JP, Rougeron A, Morio F, et al. Matrix-assisted laser desorption ionization-time of flight mass spectrometry for fast and accurate identification of Pseudallescheria/Scedosporium species. Clin Microbiol Infect. 2014; 20: 929-935. [CrossRef]
- Huprikar S, Shoham S, AST Infectious Diseases Community of Practice. Emerging fungal infections in solid organ transplantation. Am J Transplant. 2013; 13 Suppl 4: 262-271. [CrossRef]
- Husain S, Silveira FP, Azie N, Franks B, Horn D. Epidemiological features of invasive mold infections among solid organ transplant recipients: PATH Alliance® registry analysis. Med Mycol. 2017; 55: 269-277.
- Fischer SA, Lu K, AST Infectious Diseases Community of Practice. Screening of donor and recipient in solid organ transplantation. Am J Transplant. 2013; 13 Suppl 4: 9-21. [CrossRef]
- Blair JE, Ampel NM, Hoover SE. Coccidioidomycosis in selected immunosuppressed hosts. Med Mycol. 2018; 1111: 245. [CrossRef]
- Nelson JK, Giraldeau G, Montoya JG, Deresinski S, Ho DY, Pham M. Donor-derived coccidioides immitis endocarditis and disseminated infection in the setting of solid organ transplantation. Open Forum Infect Dis. 2016; 3: ofw086. [CrossRef]
- Miller R, Assi M, AST Infectious Diseases Community of Practice. Endemic fungal infections in solid organ transplantation. Am J Transplant. 2013; 13 Suppl 4: 250-261. [CrossRef]
- Singh N, Huprikar S, Burdette SD, Morris MI, Blair JE, Wheat LJ, et al. Donor-derived fungal infections in organ transplant recipients: guidelines of the American Society of Transplantation, infectious diseases community of practice. Am J Transplant. 2012; 12: 2414-2428. [CrossRef]
- Clark NM, Weigt SS, Fishbein MC, Kubak B, Belperio JA, Lynch JP. Fungal infections complicating lung transplantation. Semin Respir Crit Care Med. 2018; 39: 227-254. [CrossRef]
- Fishman JA. Infection in solid-organ transplant recipients. N Engl J Med. 2007; 357: 2601-2614. [CrossRef]
- Suzuki Y, Kenjo A, Togano T, Yamamoto N, Ohto H, Kume H. Infectious diseases in solid organ transplant recipients: Analysis of autopsied cases in Japan. J Infect Chemother. 2017; 23: 531-537. [CrossRef]
- Peleg AY, Husain S, Kwak EJ, Silveira FP, Ndirangu M, Tran J, et al. Opportunistic infections in 547 organ transplant recipients receiving alemtuzumab, a humanized monoclonal CD-52 antibody. Clin Infect Dis. 2007; 44: 204-212. [CrossRef]
- Husain S, Singh N. The impact of novel immunosuppressive agents on infections in organ transplant recipients and the interactions of these agents with antimicrobials. Clin Infect Dis. 2002; 35: 53-61. [CrossRef]
- de Boer MGJ, Bruijnesteijn van Coppenraet LES, Gaasbeek A, Berger SP, Gelinck LBS, van Houwelingen HC, et al. An outbreak of Pneumocystis jiroveci pneumonia with 1 predominant genotype among renal transplant recipients: interhuman transmission or a common environmental source? Clin Infect Dis. 2007; 44: 1143-1149. [CrossRef]
- Azar MM, Turbett SE, Fishman JA, Pierce VM. Donor-Derived Transmission of Candida auris During Lung Transplantation. Clin Infect Dis. 2017; 65: 1040-1042. [CrossRef]
- Lim C, Seo JB, Park S-Y, Hwang H-J, Lee HJ, Lee S-O, et al. Analysis of initial and follow-up CT findings in patients with invasive pulmonary aspergillosis after solid organ transplantation. Clin Radiol. 2012; 67: 1179-1186. [CrossRef]
- Park S-Y, Kim S-H, Choi S-H, Sung H, Kim M-N, Woo JH, et al. Clinical and radiological features of invasive pulmonary aspergillosis in transplant recipients and neutropenic patients. Transpl Infect Dis. 2010; 12: 309-315. [CrossRef]
- Assi M, Martin S, Wheat LJ, Hage C, Freifeld A, Avery R, et al. Histoplasmosis after solid organ transplant. Clin Infect Dis. 2013; 57: 1542-1549. [CrossRef]
- Zarrinfar H, Mirhendi H, Makimura K, Satoh K, Khodadadi H, Paknejad O. Use of mycological, nested PCR, and real-time PCR methods on BAL fluids for detection of Aspergillus fumigatus and A. flavus in solid organ transplant recipients. Mycopathologia. 2013; 176: 377-385. [CrossRef]
- Pickering JW, Sant HW, Bowles CAP, Roberts WL, Woods GL. Evaluation of a (1->3)-beta-D-glucan assay for diagnosis of invasive fungal infections. J Clin Microbiol. 2005; 43: 5957-5962. [CrossRef]
- Pappas PG, Kauffman CA, Andes DR, Clancy CJ, Marr KA, Ostrosky-Zeichner L, et al. Clinical Practice Guideline for the Management of Candidiasis: 2016 Update by the Infectious Diseases Society of America. Vol. 62, Clinical infectious diseases : an official publication of the Infectious Diseases Society of America. 2016. pp. e1–50.
- Kathuria S, Singh PK, Sharma C, Prakash A, Masih A, Kumar A, et al. Multidrug-resistant Candida auris misidentified as Candida haemulonii: Characterization by matrix-assisted laser desorption ionization-time of flight mass spectrometry and DNA sequencing and its antifungal susceptibility profile variability by vitek 2, CLSI broth microdilution, and etest method. Warnock DW, editor. J Clin Microbiol. 5 ed. American Society for Microbiology Journals; 2015; 53: 1823-1830.
- Rañó A, Agustí C, Jimenez P, Angrill J, Benito N, Danés C, et al. Pulmonary infiltrates in non-HIV immunocompromised patients: a diagnostic approach using non-invasive and bronchoscopic procedures. Thorax. BMJ Publishing Group; 2001; 56: 379-387.
- Küpeli E, Akçay S, Ulubay G, Ozyurek BA, Ozdemirel TS, Haberal M. Diagnostic utility of flexible bronchoscopy in recipients of solid organ transplants. Transplant Proc. 2011; 43: 543-546. [CrossRef]
- Hsu JL, Kuschner WG, Paik J, Bower N, Vazquez Guillamet MC, Kothary N. The diagnostic yield of CT-guided percutaneous lung biopsy in solid organ transplant recipients. Clin Transplant. 2012; 26: 615-621. [CrossRef]
- Clancy CJ, Pappas PG, Vazquez J, Judson MA, Kontoyiannis DP, Thompson GR, et al. Detecting Infections Rapidly and Easily for Candidemia Trial, Part 2 (DIRECT2): A Prospective, Multicenter Study of the T2Candida Panel. Clin Infect Dis. 2018; 66: 1678-1686. [CrossRef]
- Rabodonirina M, Piens MA, Monier MF, Guého E, Fière D, Mojon M. Fusarium infections in immunocompromised patients: case reports and literature review. Eur J Clin Microbiol Infect Dis. 1994; 13: 152-161. [CrossRef]
- Muhammed M, Anagnostou T, Desalermos A, Kourkoumpetis TK, Carneiro HA, Glavis-Bloom J, et al. Fusarium infection: report of 26 cases and review of 97 cases from the literature. Medicine (Baltimore). 2013; 92: 305-316. [CrossRef]
- Kirby JE, Delaney M, Qian Q, Gold HS. Optimal use of Myco/F lytic and standard BACTEC blood culture bottles for detection of yeast and mycobacteria. Arch Pathol Lab Med. 2009; 133: 93-96.
- Ramanan P, Vetter E, Milone AA, Patel R, Wengenack NL. Comparison of BACTEC MYCO/F Lytic Bottle to the Wampole Isolator for Recovery of Fungal and Mycobacterial Organisms. Open Forum Infect Dis. 2016; 3(suppl_1).
- Del Chierico F, Masotti A, Onori M, Fiscarelli E, Mancinelli L, Ricciotti G, et al. MALDI-TOF MS proteomic phenotyping of filamentous and other fungi from clinical origin. J Proteomics. 2012; 75: 3314-3330. [CrossRef]
- Rychert J, Slechta ES, Barker AP, Miranda E, Babady NE, Tang Y-W, et al. Multicenter Evaluation of the Vitek MS v3.0 System for the Identification of Filamentous Fungi. Warnock DW, editor. J Clin Microbiol. American Society for Microbiology Journals; 2018; 56: 1571.
- Sanguinetti M, Posteraro B. Identification of Molds by Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry. Kraft CS, editor. J Clin Microbiol. 2017; 55: 369-379. [CrossRef]
- Ciardo DE, Lucke K, Imhof A, Bloemberg GV, Böttger EC. Systematic internal transcribed spacer sequence analysis for identification of clinical mold isolates in diagnostic mycology: a 5-year study. J Clin Microbiol. 2010; 48: 2809-2813. [CrossRef]
- Guarner J, Brandt ME. Histopathologic diagnosis of fungal infections in the 21st century. Clin Microbiol Rev. 1st ed. 2011; 24: 247-280.
- Leeflang MMG, Debets-Ossenkopp YJ, Wang J, Visser CE, Scholten RJPM, Hooft L, et al. Galactomannan detection for invasive aspergillosis in immunocompromised patients. Cochrane Airways Group, editor. Cochrane Database Syst Rev. John Wiley & Sons, Ltd; 2015; 31: CD007394. [CrossRef]
- Er Dedekarginoglu B, Savas Bozbas S, Ulubay G, Oner Eyuboglu F, Haberal M. Culture-Positive Pulmonary Aspergillosis Infection: Clinical and Laboratory Features of Solid-Organ Transplant Recipients. Exp Clin Transplant. 2017; 15: 214-218.
- Pfeiffer CD, Fine JP, Safdar N. Diagnosis of invasive aspergillosis using a galactomannan assay: a meta-analysis. Clin Infect Dis. 2006; 42: 1417-1427. [CrossRef]
- Tabarsi P, Soraghi A, Marjani M, Zandian P, Baghaei P, Najafizadeh K, et al. Comparison of serum and bronchoalveolar lavage galactomannan in diagnosing invasive aspergillosis in solid-organ transplant recipients. Exp Clin Transplant. 2012; 10: 278-281. [CrossRef]
- Connolly PA, Durkin MM, Lemonte AM, Hackett EJ, Wheat LJ. Detection of histoplasma antigen by a quantitative enzyme immunoassay. Clin Vaccine Immunol. 2007; 14: 1587-1591. [CrossRef]
- Vergidis P, Walker RC, Kaul DR, Kauffman CA, Freifeld AG, Slagle DC, et al. False-positive Aspergillus galactomannan assay in solid organ transplant recipients with histoplasmosis. Transpl Infect Dis. 2012; 14: 213-217. [CrossRef]
- Vergidis P, Razonable RR, Wheat LJ, Estes L, Caliendo AM, Baden LR, et al. Reduction in false-positive Aspergillus serum galactomannan enzyme immunoassay results associated with use of piperacillin-tazobactam in the United States. J Clin Microbiol. 2014; 52: 2199-2201. [CrossRef]
- Marr KA, Laverdiere M, Gugel A, Leisenring W. Antifungal therapy decreases sensitivity of the Aspergillus galactomannan enzyme immunoassay. Clin Infect Dis. 2005; 40: 1762-1769. [CrossRef]
- Patterson TF, Thompson GR, Denning DW, Fishman JA, Hadley S, Herbrecht R, et al. Executive Summary: Practice Guidelines for the Diagnosis and Management of Aspergillosis: 2016 Update by the Infectious Diseases Society of America. Clin Infect Dis. 2016; 63: 433-442. [CrossRef]
- Onishi A, Sugiyama D, Kogata Y, Saegusa J, Sugimoto T, Kawano S, et al. Diagnostic accuracy of serum 1,3-β-D-glucan for pneumocystis jiroveci pneumonia, invasive candidiasis, and invasive aspergillosis: systematic review and meta-analysis. J Clin Microbiol. 2012; 50: 7-15. [CrossRef]
- Slotkin R, Abi-Raad R, Liu Y, Grant M, Malinis M, Azar MM. Diagnosis of Pneumocystis jiroveci pneumonia in HIV-negative immunocompromised patients: Is the Gomori-methenamine silver stain of bronchoalveolar lavage fluid the gold standard or sub-standard? San Fransisco, CA; 2018.
- Alexander BD, Smith PB, Davis RD, Perfect JR, Reller LB. The (1,3){beta}-D-glucan test as an aid to early diagnosis of invasive fungal infections following lung transplantation. J Clin Microbiol. 2010; 48: 4083-4088. [CrossRef]
- Mutschlechner W, Risslegger B, Willinger B, Hoenigl M, Bucher B, Eschertzhuber S, et al. Bronchoalveolar Lavage Fluid (1,3)β-D-Glucan for the Diagnosis of Invasive Fungal Infections in Solid Organ Transplantation: A Prospective Multicenter Study. Transplantation. 2015; 99: e140-e144. [CrossRef]
- Chai LYA, Kullberg B-J, Johnson EM, Teerenstra S, Khin LW, Vonk AG, et al. Early serum galactomannan trend as a predictor of outcome of invasive aspergillosis. J Clin Microbiol. 2012; 50: 2330-2336. [CrossRef]
- Lyons JL, Roos KL, Marr KA, Neumann H, Trivedi JB, Kimbrough DJ, et al. Cerebrospinal fluid (1,3)-β-D-glucan detection as an aid for diagnosis of iatrogenic fungal meningitis. J Clin Microbiol. American Society for Microbiology Journals; 2013; 51: 1285-1287.
- Azar MM, Hage CA. Laboratory Diagnostics for Histoplasmosis. Kraft CS, editor. J Clin Microbiol. 2017; 55: 1612-1620. [CrossRef]
- Hage CA, Ribes JA, Wengenack NL, Baddour LM, Assi M, McKinsey DS, et al. A multicenter evaluation of tests for diagnosis of histoplasmosis. Clin Infect Dis. 2011; 53: 448-454. [CrossRef]
- Theel ES, Ramanan P. Clinical Significance of Low-Positive Histoplasma Urine Antigen Results. Warnock DW, editor. J Clin Microbiol. 2014; 52: 3444-3446. [CrossRef]
- Wheat J, Myint T, Guo Y, Kemmer P, Hage C, Terry C, et al. Central nervous system histoplasmosis: Multicenter retrospective study on clinical features, diagnostic approach and outcome of treatment. Medicine (Baltimore). 2018; 97: e0245. [CrossRef]
- Durkin M, Witt J, Lemonte A, Wheat B, Connolly P. Antigen assay with the potential to aid in diagnosis of blastomycosis. J Clin Microbiol. American Society for Microbiology Journals; 2004; 42: 4873-4875.
- Gade W, Hinnefeld SW, Babcock LS, Gilligan P, Kelly W, Wait K, et al. Comparison of the PREMIER cryptococcal antigen enzyme immunoassay and the latex agglutination assay for detection of cryptococcal antigens. J Clin Microbiol. 1991; 29: 1616-1619.
- Menichetti F, Fiorio M, Tosti A, Gatti G, Bruna Pasticci M, Miletich F, et al. High-dose fluconazole therapy for cryptococcal meningitis in patients with AIDS. Clin Infect Dis. 1996; 22: 838-840. [CrossRef]
- Osawa R, Alexander BD, Lortholary O, Dromer F, Forrest GN, Lyon GM, et al. Identifying predictors of central nervous system disease in solid organ transplant recipients with cryptococcosis. Transplantation. 2010; 89: 69-74. [CrossRef]
- Mendoza N, Blair JE. The utility of diagnostic testing for active coccidioidomycosis in solid organ transplant recipients. Am J Transplant. 2013; 13: 1034-1039. [CrossRef]
- Haselow D, Fields VS, Fialkowski V, Gibbons-Brugener S, Jackson B, Pedati C, et al. Standardized surveillance case definition for histoplasmosis. https://cdn.ymaws.com/www.cste.org/resource/resmgr/2016PS/16_ID_02.pdf. 2016.
- Powers-Fletcher MV, Hanson KE. Nonculture Diagnostics in Fungal Disease. Infect Dis Clin North Am. 2016; 30: 37-49. [CrossRef]
- Richer SM, Smedema ML, Durkin MM, Brandhorst TT, Hage CA, Connolly PA, et al. Development of a highly sensitive and specific blastomycosis antibody enzyme immunoassay using Blastomyces dermatitidis surface protein BAD-1. Papasian CJ, editor. Clin Vaccine Immunol. American Society for Microbiology; 2014; 21: 143-146.
- Avni T, Leibovici L, Paul M. PCR Diagnosis of Invasive Candidiasis: Systematic Review and Meta-Analysis. J Clin Microbiol. 2011; 49: 665-670. [CrossRef]
- Powers-Fletcher MV, Hanson KE. Molecular Diagnostic Testing for Aspergillus. Kraft CS, editor. J Clin Microbiol. 2016; 54: 2655-2660. [CrossRef]
- White PL, Bretagne S, Klingspor L, Melchers WJG, McCulloch E, Schulz B, et al. Aspergillus PCR: one step closer to standardization. J Clin Microbiol. 2010; 48: 1231-1240. [CrossRef]
- Cruciani M, Mengoli C, Loeffler J, Donnelly P, Barnes R, Jones BL, et al. Polymerase chain reaction blood tests for the diagnosis of invasive aspergillosis in immunocompromised people. Cochrane Gynaecological, Neuro-oncology and Orphan Cancer Group, editor. Cochrane Database Syst Rev. 2015; 4: CD009551. [CrossRef]
- Imbert S, Brossas J-Y, Palous M, Joly I, Meyer I, Fekkar A. Performance of Aspergillus PCR in cerebrospinal fluid for the diagnosis of cerebral aspergillosis. Clin Microbiol Infect. 2017; 23: 889.e1-889.e4. [CrossRef]
- Hauser PM, Bille J, Lass-Flörl C, Geltner C, Feldmesser M, Levi M, et al. Multicenter, prospective clinical evaluation of respiratory samples from subjects at risk for Pneumocystis jirovecii infection by use of a commercial real-time PCR assay. J Clin Microbiol. 2011; 49: 1872-1878. [CrossRef]
- Slechta ES, Hohmann SL, Simmon K, Hanson KE. Internal transcribed spacer region sequence analysis using SmartGene IDNS software for the identification of unusual clinical yeast isolates. Med Mycol. 2012; 50: 458-466. [CrossRef]
- Lau A, Sorrell TC, Chen S, Stanley K, Iredell J, Halliday C. Multiplex tandem PCR: a novel platform for rapid detection and identification of fungal pathogens from blood culture specimens. J Clin Microbiol. 2008; 46: 3021-3027. [CrossRef]
- Otlu B, Bayindir Y, Ozdemir F, Ince V, Cuglan S, Hopoglu M, et al. Rapid Detection of Bloodstream Pathogens in Liver Transplantation Patients With FilmArray Multiplex Polymerase Chain Reaction Assays: Comparison With Conventional Methods. Transplant Proc. 2015; 47: 1926-1932. [CrossRef]
- Salimnia H, Fairfax MR, Lephart PR, Schreckenberger P, DesJarlais SM, Johnson JK, et al. Evaluation of the FilmArray Blood Culture Identification Panel: Results of a Multicenter Controlled Trial. Richter SS, editor. J Clin Microbiol. 9 ed. American Society for Microbiology Journals; 2016; 54: 687-698.
- Altun O, Almuhayawi M, Ullberg M, Ozenci V. Clinical evaluation of the FilmArray blood culture identification panel in identification of bacteria and yeasts from positive blood culture bottles. J Clin Microbiol. 2013; 51: 4130-4136. [CrossRef]
- Hanson KE, Slechta ES, Killpack JA, Heyrend C, Lunt T, Daly JA, et al. Preclinical Assessment of a Fully Automated Multiplex PCR Panel for Detection of Central Nervous System Pathogens. Richter SS, editor. J Clin Microbiol. 2016; 54: 785-787. [CrossRef]
- Mylonakis E, Clancy CJ, Ostrosky-Zeichner L, Garey KW, Alangaden GJ, Vazquez JA, et al. T2 magnetic resonance assay for the rapid diagnosis of candidemia in whole blood: a clinical trial. Clin Infect Dis. 2015; 60: 892-899. [CrossRef]
- Clancy CJ, Nguyen MH. Finding the “missing 50%” of invasive candidiasis: how nonculture diagnostics will improve understanding of disease spectrum and transform patient care. Clin Infect Dis. 2013; 56: 1284-1292. [CrossRef]
- Muñoz P, Vena A, Machado M, Martínez-Jiménez MC, Gioia F, Gómez E, et al. T2MR contributes to the very early diagnosis of complicated candidaemia. A prospective study. J Antimicrob Chemother. 2018; 73: iv13-iv19. [CrossRef]
- Muñoz P, Vena A, Machado M, Gioia F, Martínez-Jiménez MC, Gómez E, et al. T2Candida MR as a predictor of outcome in patients with suspected invasive candidiasis starting empirical antifungal treatment: a prospective pilot study. J Antimicrob Chemother. 2018; 73: iv6–iv12. [CrossRef]
- Giannella M, Paolucci M, Roncarati G, Vandi G, Pascale R, Trapani F, et al. Potential role of T2Candida in the management of empirical antifungal treatment in patients at high risk of candidaemia: a pilot single-centre study. J Antimicrob Chemother. 2018; 73: 2856-2859. [CrossRef]
- Walker B, Powers-Fletcher MV, Schmidt RL, Hanson KE. Cost-Effectiveness Analysis of Multiplex PCR with Magnetic Resonance Detection versus Empiric or Blood Culture-Directed Therapy for Management of Suspected Candidemia. Warnock DW, editor. J Clin Microbiol. 3rd ed. 2016; 54: 718-726. [CrossRef]
- Bilir SP, Ferrufino CP, Pfaller MA, Munakata J. The economic impact of rapid Candida species identification by T2Candida among high-risk patients. Future Microbiol. 2015; 10: 1133-1144. [CrossRef]
- Pfeiffer CD, Samsa GP, Schell WA, Reller LB, Perfect JR, Alexander BD. Quantitation of Candida CFU in initial positive blood cultures. J Clin Microbiol. 2011; 49: 2879-2883. [CrossRef]
- Schlaberg R, Queen K, Simmon K, Tardif K, Stockmann C, Flygare S, et al. Viral Pathogen Detection by Metagenomics and Pan-Viral Group Polymerase Chain Reaction in Children With Pneumonia Lacking Identifiable Etiology. J Infect Dis. 2017; 215: 1407-1415. [CrossRef]
- Charpentier E, Garnaud C, Wintenberger C, Bailly S, Murat J-B, Rendu J, et al. Added Value of Next-Generation Sequencing for Multilocus Sequence Typing Analysis of a Pneumocystis jirovecii Pneumonia Outbreak1. Emerging Infect Dis. 2017; 23: 1237-1245. [CrossRef]
- Zhang Y, Ai J-W, Cui P, Wu H-L, Ye M-Z, Zhang W-H. A cluster of cases of pneumocystis pneumonia identified by shotgun metagenomics approach. J Infect. 2018. [CrossRef]
- Zinter MS, Dvorak CC, Mayday MY, Iwanaga K, Ly NP, McGarry ME, et al. Pulmonary Metagenomic Sequencing Suggests Missed Infections in Immunocompromised Children. Clin Infect Dis. 2018. [CrossRef]
- Hayden RT, Sun Y, Tang L, Procop GW, Hillyard DR, Pinsky BA, et al. Progress in Quantitative Viral Load Testing: Variability and Impact of the WHO Quantitative International Standards. McAdam AJ, editor. J Clin Microbiol. 2017; 55: 423-430. [CrossRef]
- Morrissey CO, Chen SC-A, Sorrell TC, Milliken S, Bardy PG, Bradstock KF, et al. Galactomannan and PCR versus culture and histology for directing use of antifungal treatment for invasive aspergillosis in high-risk haematology patients: a randomised controlled trial. Lancet Infect Dis. 2013; 13: 519-528. [CrossRef]
- Chryssanthou E, Klingspor L, Tollemar J, Petrini B, Larsson L, Christensson B, et al. PCR and other non-culture methods for diagnosis of invasive Candida infections in allogeneic bone marrow and solid organ transplant recipients. Mycoses. 1999; 42: 239-247. [CrossRef]
- Muller N, Kessler R, Caillard S, Epailly E, Hubelé F, Heimburger C, et al. 18F-FDG PET/CT for the Diagnosis of Malignant and Infectious Complications After Solid Organ Transplantation. Nucl Med Mol Imaging. 2017; 51: 58-68. [CrossRef]
- Hot A, Maunoury C, Poiree S, Lanternier F, Viard JP, Loulergue P, et al. Diagnostic contribution of positron emission tomography with [18F]fluorodeoxyglucose for invasive fungal infections. Clin Microbiol Infect. 2011; 17: 409-417. [CrossRef]
- Egli A, Osthoff M, Goldenberger D, Halter J, Schaub S, Steiger J, et al. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF) directly from positive blood culture flasks allows rapid identification of bloodstream infections in immunosuppressed hosts. Transpl Infect Dis. 2015; 17: 481-487. [CrossRef]
- Radic M, Goic-Barisic I, Novak A, Rubic Z, Tonkic M. Evaluation of PNA FISH® Yeast Traffic Light in identification of Candida species from blood and non-blood culture specimens. Med Mycol. 2016; 54: 654-658. [CrossRef]
- Gorton RL, Ramnarain P, Barker K, Stone N, Rattenbury S, McHugh TD, et al. Comparative analysis of Gram's stain, PNA-FISH and Sepsityper with MALDI-TOF MS for the identification of yeast direct from positive blood cultures. Mycoses. 7 ed. 2014; 57: 592-601. [CrossRef]
- Hanson KE, Couturier MR. Multiplexed Molecular Diagnostics for Respiratory, Gastrointestinal, and Central Nervous System Infections. Barth Reller L, Weinstein MP, editors. Clin Infect Dis. 2016; 63: 1361-1367.
- Leber AL, Everhart K, Balada-Llasat J-M, Cullison J, Daly J, Holt S, et al. Multicenter Evaluation of BioFire FilmArray Meningitis/Encephalitis Panel for Detection of Bacteria, Viruses, and Yeast in Cerebrospinal Fluid Specimens. Forbes BA, editor. J Clin Microbiol. 2016; 54: 2251-2261. [CrossRef]
- Valero C, la Cruz-Villar de L, Zaragoza Ó, Buitrago MJ. New Panfungal Real-Time PCR Assay for Diagnosis of Invasive Fungal Infections. J Clin Microbiol. 2016; 54: 2910-2918. [CrossRef]
- Capoor MR, Puri S, Raheja H, Mohindra R, Gupta DK, Verma PK, et al. Screening of invasive fungal infections by a real-time panfungal (pan-ACF) polymerase chain reaction assay in patients with haematological malignancy. Indian J Med Microbiol. 2017; 35: 41-47. [CrossRef]
- Micó M, Navarro F, de Miniac D, González Y, Brell A, López C, et al. Efficacy of the FilmArray blood culture identification panel for direct molecular diagnosis of infectious diseases from samples other than blood. J Med Microbiol. 2015; 64: 1481-1488. [CrossRef]
- Damiani C, Le Gal S, Da Costa C, Virmaux M, Nevez G, Totet A. Combined quantification of pulmonary Pneumocystis jirovecii DNA and serum (1->3)-β-D-glucan for differential diagnosis of pneumocystis pneumonia and Pneumocystis colonization. J Clin Microbiol. 2013; 51: 3380-3388. [CrossRef]
- Moncada PA, Budvytiene I, Ho DY, Deresinski SC, Montoya JG, Banaei N. Utility of DNA sequencing for direct identification of invasive fungi from fresh and formalin-fixed specimens. Am J Clin Pathol. 2013; 140: 203-208. [CrossRef]
- Koo S, Thomas HR, Daniels SD, Lynch RC, Fortier SM, Shea MM, et al. A breath fungal secondary metabolite signature to diagnose invasive aspergillosis. Clin Infect Dis. 2014; 59: 1733-1740. [CrossRef]
- Willinger B, Lackner M, Lass-Flörl C, Prattes J, Posch V, Selitsch B, et al. Bronchoalveolar lavage lateral-flow device test for invasive pulmonary aspergillosis in solid organ transplant patients: a semiprospective multicenter study. Transplantation. 2014; 98: 898-902. [CrossRef]
- Florescu DF, Sandkovsky U. Fungal infections in intestinal and multivisceral transplant recipients. Curr Opin Organ Transplant. 2015; 20: 295-302. [CrossRef]


